
TLDR: Earlier we showed how LLM agents can merge large tables with no shared key. Here, we show that asking the agents to classify the problem type first (1:1, 1:many, many:1), and then do the matching, we get a much reduced false positve rate.
LLM agents can use judgment and web research to do hard semantic data tasks, like matching a table of CEOs to a table of companies. But we sometimes saw high false positive rates, e.g. LLM would confidently assign multiple CEOs to the same company, because each individual match looked plausible even though the aggregate result was clearly wrong.
What if we had the agent classify the problem type first? LLMs should be able to judge whether a matching should be 1:1, 1:many, many:1, or many:many as well as a human. We tested how this worked on a bunch of thorny table merging problems.
1. One-to-One
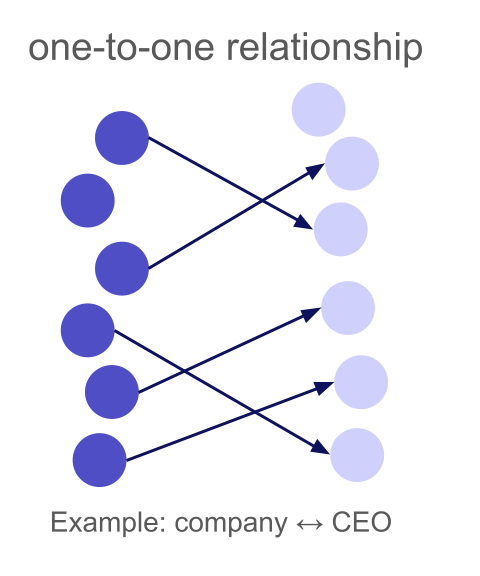
A one-to-one relationship means each left row matches at most one right row, and each right row matches at most one left row. This is the most constrained type: a bijection between matched subsets.
To illustrate this, let us look again at our example (Experiment 3) from the previous article where we merged 438 companies to their CEOs (we implicitly used a many-to-one relationship type). The LLM (Gemini-3-Flash low reasoning, which is our default model for merge) did well on well-known executives (like Tim Cook and Satya Nadella), but for less famous CEOs, it sometimes assigned the same person to multiple companies. In total, it only had an accuracy of 87.3% with 12.3% false positives.
Now let us add the one-to-one constraint to the same table merge:
result = await merge(
task="Merge the CEO to the company information, use web search if unsure",
left_table=left_table,
right_table=right_table,
relationship_type="one_to_one"
)
| relationship type | Matched | LLM Match | Web Match | Accuracy | False Positives | Cost |
|---|---|---|---|---|---|---|
| many-to-one | 100% | 98.9% | 1.1% | 87.7% | 12.3% | $0.75 |
| one-to-one | 98.4% | 77.9% | 20.5% | 96.3% | 2.1% | $1.50 |
With one_to_one, clashed rows are re-run with web verification. Some genuinely ambiguous cases are left unmatched (the 1.6% drop in match rate), but false positives drop from 12.3% to 2.1%. Looking at the false positives, these happened from the LLM merge without web search, so they did not lead to clashes, i.e. the LLM was just too confident here. All clashes have been resolved and correctly matched.
The cost increase comes from web search re-runs on clashed rows. We pay about $0.75 extra to eliminate almost all false positives from a 438 row merge.
2. Many-to-One
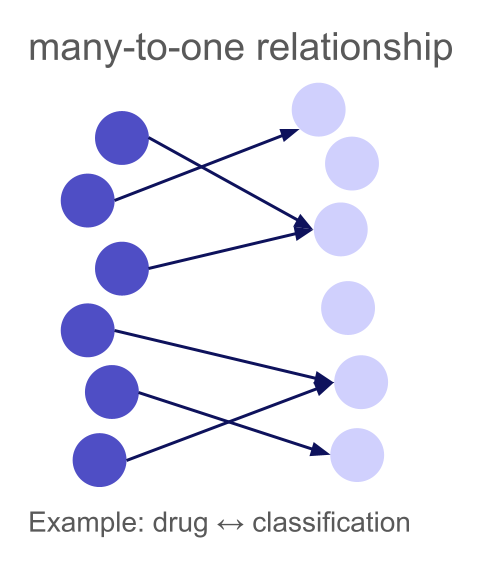
A many-to-one relationship means multiple left rows can match the same right row, but each left row matches at most one right row. This is common when mapping entities to categories, taxonomies, or finding the "parent" of each item.
Drugs to Classifications: When the LLM Already Knows
A natural many-to-one example is mapping entities to a fixed taxonomy. We have 100 common drugs and 39 ATC (Anatomical Therapeutic Chemical) classifications:
Left table (300 drugs):
| drug |
|---|
| Paracetamol |
| Ibuprofen |
| Morphine |
| Amoxicillin |
| ... |
Right table (133 classifications):
| atc_classification |
|---|
| Analgesic |
| NSAID |
| Opioid analgesic |
| Penicillin antibiotic |
| ... |
Multiple drugs share the same classification (Ibuprofen, Naproxen, and Diclofenac are all NSAIDs), but each drug has exactly one primary ATC classification.
result = await merge(
task="Find the ATC classification of each drug",
left_table=drugs_df,
right_table=classifications_df,
relationship_type="many_to_one",
)
| Matched | LLM Match | Web Match | Accuracy | False Positives | Cost |
|---|---|---|---|---|---|
| 100% | 100% | 0% | 98.7% | 1.3% | $0.44 |
Drug classifications are well-established medical knowledge, so LLMs know about them, so no need for web search. The many_to_one constraint is important here: it tells the system that it's perfectly fine for multiple drugs to map to "NSAID" or "Opioid analgesic". Without this, the system might flag these shared mappings as clashes and waste time re-running them.
Looking at the false positives, these were really edge cases, like Sotalol being classified as "Beta blocker" instead of "Class III antiarrhythmic", one of the very few drugs that actually has two classifications because it is used to treat multiple conditions.
Not all many-to-one merges are this straightforward. Consider matching research papers to their first (main) author from a list of researchers. Each paper has exactly one first author, but a researcher is typically the first author on several papers in our dataset — a classic many-to-one relationship.
Left table (895 papers):
| paper |
|---|
| Approaching Human-Level Forecasting with Language Models |
| Adaptive social networks promote the wisdom of crowds |
| When combinations of humans and AI are useful |
| ... |
Right table (303 researchers):
| name |
|---|
| Danny Halawi |
| Abdullah Almaatouq |
| Houtan Bastani |
| ... |
result = await merge(
task="Merge first authors to research papers, only match when you are very sure, otherwise rely on supporting web-search",
left_table=papers_df, # 895 papers
right_table=authors_df, # 303 researchers
relationship_type="many_to_one",
use_web_search=True,
)
As you see, we have been very explicit in the prompt that the LLM should only match if it is confident to reduce hallucinations. Also, we are forcing web-search for every row, as this task is almost impossible for the LLM to solve without it.
| Matched | LLM Match | Web Match | Accuracy | False Positives | Cost |
|---|---|---|---|---|---|
| 97.6% | 0% | 97.6% | 97% | 0.6% | $7.62 |
Finding the first author is a straightforward lookup. A single web search for the paper title reveals the author list immediately.
3. One-to-Many
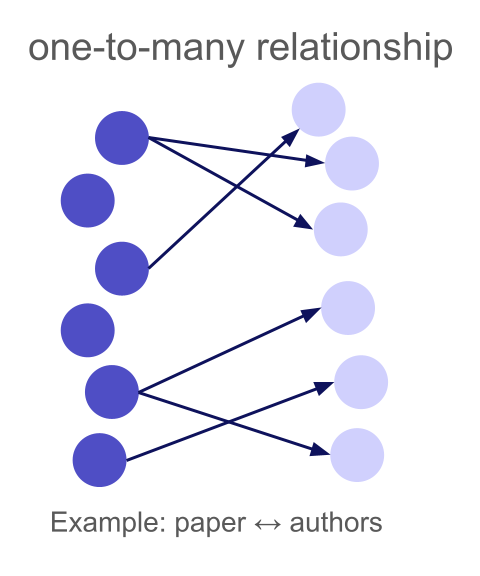
A one-to-many relationship means each left row can match multiple right rows, but each right row matches at most one left row. The "one" side is the left table, the "many" side is the right.
"Isn't one-to-many just many-to-one with the tables swapped?" Technically, yes, but table order matters enormously for agent-based merging. Wrong direction can tank both accuracy and cost.
Our merge strategy processes one left row at a time. For each left row, it shows the LLM (or web agent) that single row alongside the entire right table (or a batch of it if the table is too large) and asks: "which right row(s) match?" The web agent also researches the left row specifically, looking up information to help disambiguate.
This means:
- The left table should contain the entities you want to research individually
- The right table should contain the candidates you're matching against
But there are more reasons why table order matters. In our setup, where we match one entity from the left table with the right table, it is basically a "needle in the haystack" problem: the LLM has to pay attention to all entities of the right table at once and find the right pattern in it. And of course this works better if the "haystack" is rather small. So if you for example match a huge table of 5000 drugs to just 50 classifications, it is very easy for the LLM to find the right classification for each drug. But when you look for all drugs for a specific classification, the LLM always has to split its attention to all drugs at once and pick the right ones. In the end this is a tradeoff between number of agents and context length:
- Left table large, right table small: many agents but little context per agent
- Left table small, right table large: few agents but lots of context per agent
So depending on how hard the matching is, you can utilize more agents to trade accuracy for resources.
Reversing table order
To illustrate why direction matters, let's reverse the paper-author merge from section 2. Instead of "find the first author of this paper" (many_to_one), we now ask "find all first authored papers of this researcher from this list" (one_to_many).
We swap the dataframes, swap the relationship type, and adapt the prompt:
# Reversed: researchers on the left, papers on the right
result = await merge(
task="For each researcher, find the papers from the list where this researcher is the first author. Only match if you see evidence in the supporting web search that the researcher is really the first author of the respective paper.",
left_table=authors_df, # 303 researchers
right_table=papers_df, # 895 papers
relationship_type="one_to_many",
use_web_search=True,
)
Again, we have been very explicit about requiring web search and first author evidence. However, the results now look quite different:
| Configuration | Matched | LLM Match | Web Match | Accuracy | False Positives | Cost |
|---|---|---|---|---|---|---|
Papers → authors, many_to_one | 97.6% | 0% | 97.6% | 97% | 0.6% | $7.62 |
Authors → papers, one_to_many | 95.7% | 0% | 95.7% | 83.3% | 40.4% | $32.75 |
The reversed configuration is worse on every metric. The false positive rate is enormous despite the clash detection and resolution. This is because finding "which papers did this researcher first author?" is fundamentally harder than "who is the first author of this paper?" A paper's author list is right there on the abstract page — one web search, one answer. But a prolific researcher may have published hundreds of papers, and the agent has to figure out which specific papers from our list of 895 they were the first author on. That requires navigating Google Scholar profiles, university pages, or DBLP, and then cross referencing against a long list of candidate papers. That also explains the cost difference: finding the authors of a paper is a single web search, while finding the papers of a researcher needs parsing several websites.
Where one-to-many truly shines is when the "one" side contains well defined, searchable entities and the "many" side contains noisy, unidentifiable items. Consider matching IKEA product IDs to their product descriptions:
Left table (22 product IDs):
| product_id |
|---|
| 10018194 |
| 10032294 |
| 100467 |
| ... |
Right table (189 descriptions):
| product_description |
|---|
| ORDNING Dish drainer, stainless steel, 50x27x36 cm |
| An ORDNING stainless-steel dish drainer with two-tiered racks... |
| ORDNING Secaplatos, acero inoxidable, 50x27x36 cm |
| ... |
Each IKEA product has multiple descriptions (different languages, regional variants), but each description belongs to exactly one product — a natural one-to-many relationship. Product IDs are searchable (the agent can look up "IKEA 10018194" and find it's the ORDNING dish drainer), while descriptions are too unspecific to match the other way — "ORDNING Dish drainer, stainless steel" could match many products without the ID context.
result = await merge(
task="Merge the product descriptions to the IKEA product IDs.",
left_table=product_ids_df, # 22 product IDs
right_table=product_descriptions_df, # 189 descriptions
relationship_type="one_to_many",
use_web_search="yes",
)
| Matched | LLM Match | Web Match | Accuracy | False Positives | Cost |
|---|---|---|---|---|---|
| 90.9% | 0% | 90.9% | 94.2% | 0% | 0.36$ |
The matching works reasonably well here. Product IDs are well defined entities the agent can research. For example, the web agents collected information for product ID 10018194:
The IKEA product with ID 10018194 (also formatted as 100.181.94) is the ORDNING Dish Drainer.
Key details for matching:
Product Name: ORDNING
Description: Dish drainer, stainless steel
Dimensions: 19 5/8 x 10 5/8 x 14 1/8 " (50 x 27 x 36 cm)
Material: Stainless steel
Capacity: Holds up to 29 plates
Designer: Mikael Warnhammar
Descriptions, on the other hand, are short snippets in various languages: you can't meaningfully search for which product a generic description belongs to without already knowing the product. Reversing this merge (descriptions on the left, product IDs on the right, many_to_one) would therefore not work here.
4. Many-to-Many
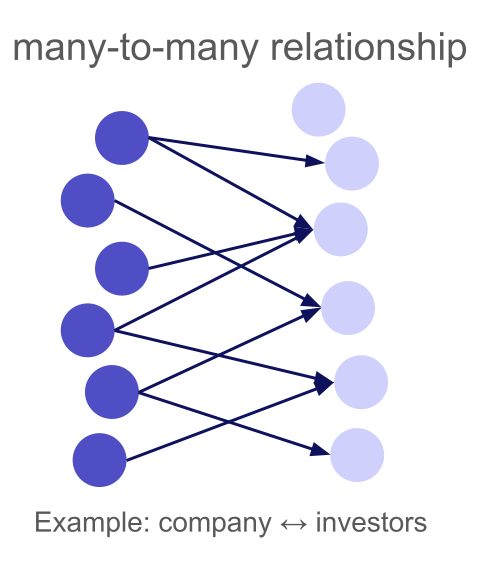
A many-to-many relationship is the most general case: any left row can match multiple right rows, and any right row can match multiple left rows. There are no uniqueness constraints at all.
Research papers and their (co-)authors are a natural example. Each paper has multiple authors, and each author has coauthored multiple papers. Unlike the many-to-one example in Section 2, where we matched papers to just their first author, here we want to find all authors of each paper from a list of researchers, making this a true many-to-many relationship.
We have 117 papers about LLM uncertainty, forecasting, and human AI collaboration, and 781 researchers:
Left table (117 papers):
| paper_name |
|---|
| Approaching Human-Level Forecasting with Language Models |
| When combinations of humans and AI are useful |
| Detecting hallucinations in large language models using semantic entropy |
| ForecastBench: A Dynamic Benchmark of AI Forecasting Capabilities |
| ... |
Right table (781 researchers):
| author |
|---|
| Danny Halawi |
| Michelle Vaccaro |
| Sebastian Farquhar |
| Yarin Gal |
| ... |
Again, we put papers on the left because looking up all authors of a specific paper is straightforward with a single web search for the paper title, in contrast to finding all publications of a researcher.
result = await merge(
task="Merge the (co-)authors / researchers to the papers",
left_table=papers_df,
right_table=authors_df,
relationship_type="many_to_many",
use_web_search="yes",
)
| Matched | LLM Match | Web Match | Accuracy | False Positives | Cost |
|---|---|---|---|---|---|
| 97.4% | 0% | 97.4% | 95.7% | 6.2% | $3.94 |
Many-to-many is the hardest relationship type. The system cannot use clash detection (any match pattern is theoretically valid), so it relies entirely on the quality of individual match decisions.
Compared to the many-to-one case in Section 2 (where we only needed the first author), the many-to-many match must identify all co-authors, which means more items to match per row and more opportunities for both misses and false positives.
Summary
| Relationship Type | Use When | Clash Detection | Table Order |
|---|---|---|---|
one_to_one | Each entity matches exactly one counterpart | Both sides | Either order works |
one_to_many | Left entity maps to multiple right entities | Right side | Put the "one" on the left |
many_to_one | Multiple left entities share a right entity | None | Put the item to research on the left |
many_to_many | No uniqueness constraints | None | Put the entity with fewer relationships on the left |
Cardinality is not just a schema annotation — it's a constraint that helps the system produce better results. By detecting and resolving clashes, one-to-one and one-to-many merges achieve significantly lower false positive rates. And by choosing the right table order for many-to-one and many-to-many, you make the web agents' job easier, improving both accuracy and cost.
Try It Yourself
All of the above examples use the relationship_type parameter in the everyrow SDK. Install it and try on your own data:
pip install everyrow
from everyrow.ops import merge
result = await merge(
task="Describe your merge task here",
left_table=your_left_df,
right_table=your_right_df,
relationship_type="one_to_one", # or "one_to_many", "many_to_one", "many_to_many"
)
Importantly, Claude can figure out which relationship_type to use! Install the everyrow plugin into Claude Code, and then when it needs to deploy a bunch of web agents to solve a problem, it can figure out these params for you.